Within families, proteins have similar tertiary structures and functional similarities, which can occur with as low as 15% sequence identity. The study of protein evolution has been energized by the genomic revolution, which greatly expanded the numbers of sequences that associate into various protein families. Software implementing conservation and co-evolution analyses is available at. 
This possibility should be included when designing and interpreting sequence analyses of other protein families. Therefore, the tertiary structure can accommodate multiple networks of functionally important positions. Results showed that conserved positions use a mixture of the “hard-wired” and “accommodating” scaffold frameworks, but that the co-evolution networks were highly dissimilar between any pair of subfamilies. Functionally important positions were identified by conservation and co-evolutionary sequence analyses. LacI/GalR paralogs share a common tertiary structure, but have low sequence identity (≤30%), and regulate a variety of metabolic processes. To discriminate between these possibilities, we compared the set of functionally important sites in the six largest paralogous subfamilies of the LacI/GalR transcription repressor family. Alternatively, the tertiary scaffold might be adaptable, accommodating a unique set of functionally important sites for each paralogous function. First, the locations of functionally-important sites might be “hard-wired” into the structure, with novel functions evolved by altering the amino acid (e.g. 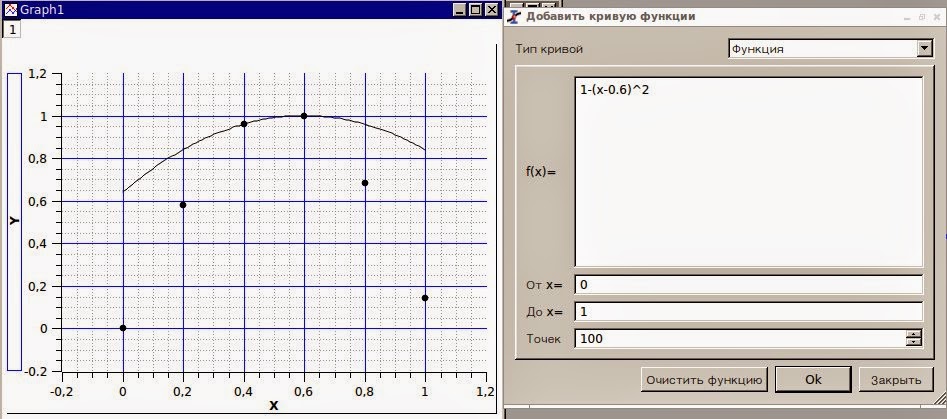
Protein families might evolve paralogous functions on their common tertiary scaffold in two ways.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
